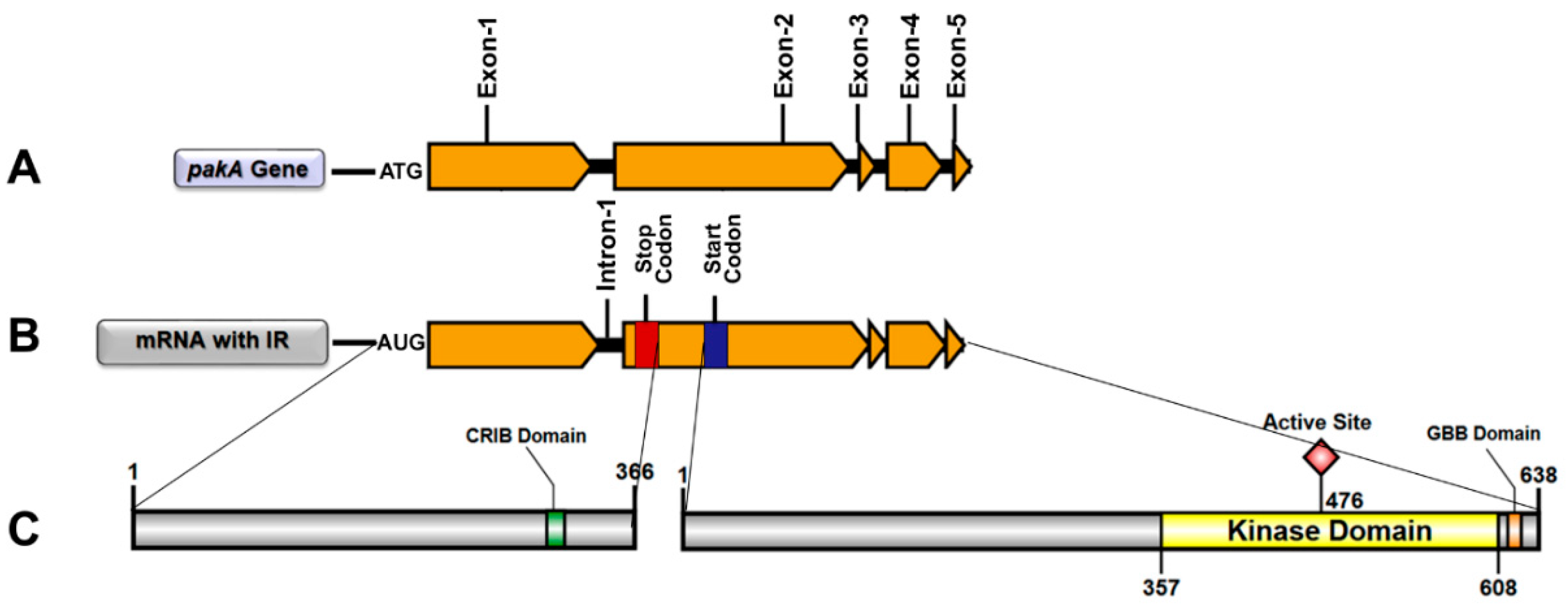
IJMS | Free Full-Text | STE20/PAKA Protein Kinase Gene Releases an Autoinhibitory Domain through Pre-mRNA Alternative Splicing in the Dermatophyte Trichophyton rubrum | HTML

PDF) Phylogenetic Comparison and Splicing Analysis of the U1 snRNP-specific Protein U1C in Eukaryotes
Spliced Leader Trapping Reveals Widespread Alternative Splicing Patterns in the Highly Dynamic Transcriptome of Trypanosoma brucei
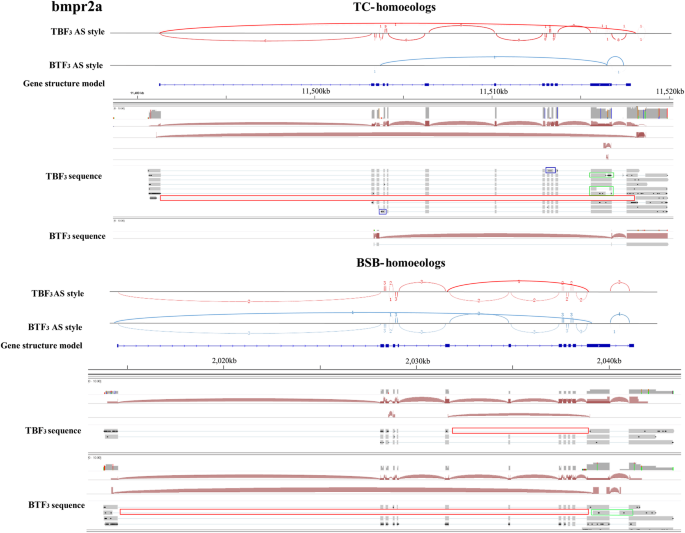
Maternal effects shape the alternative splicing of parental alleles in reciprocal cross hybrids of Megalobrama amblycephala × Culter alburnus | SpringerLink
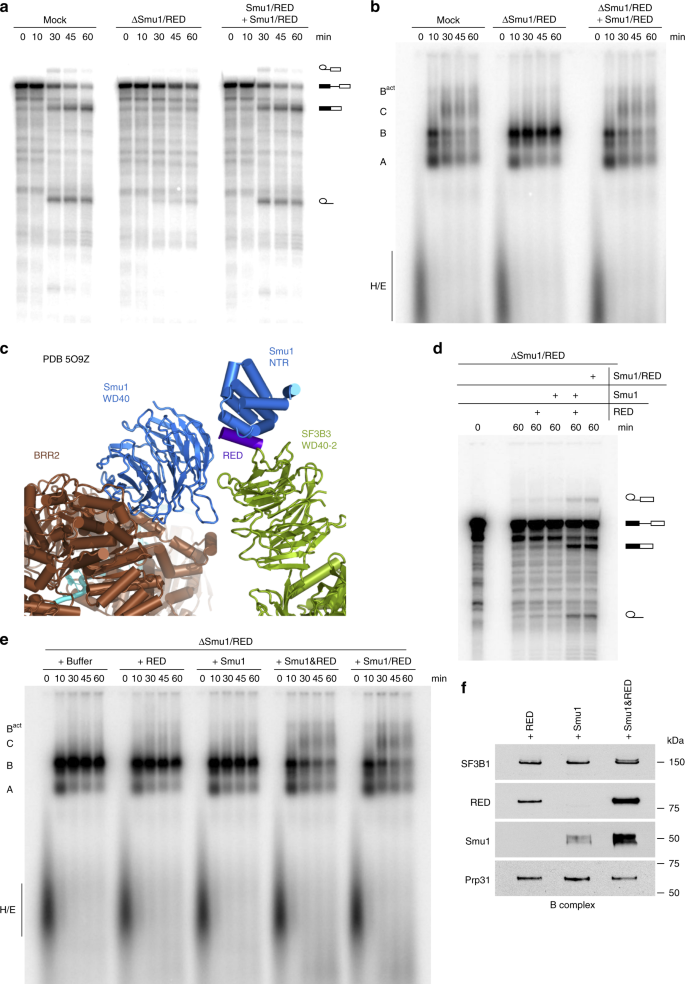
Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns | Nature Communications

PDF) Splicy: a web-based tool for the prediction of possible alternative splicing events from Affymetrix probeset data | Alessandro Guffanti - Academia.edu
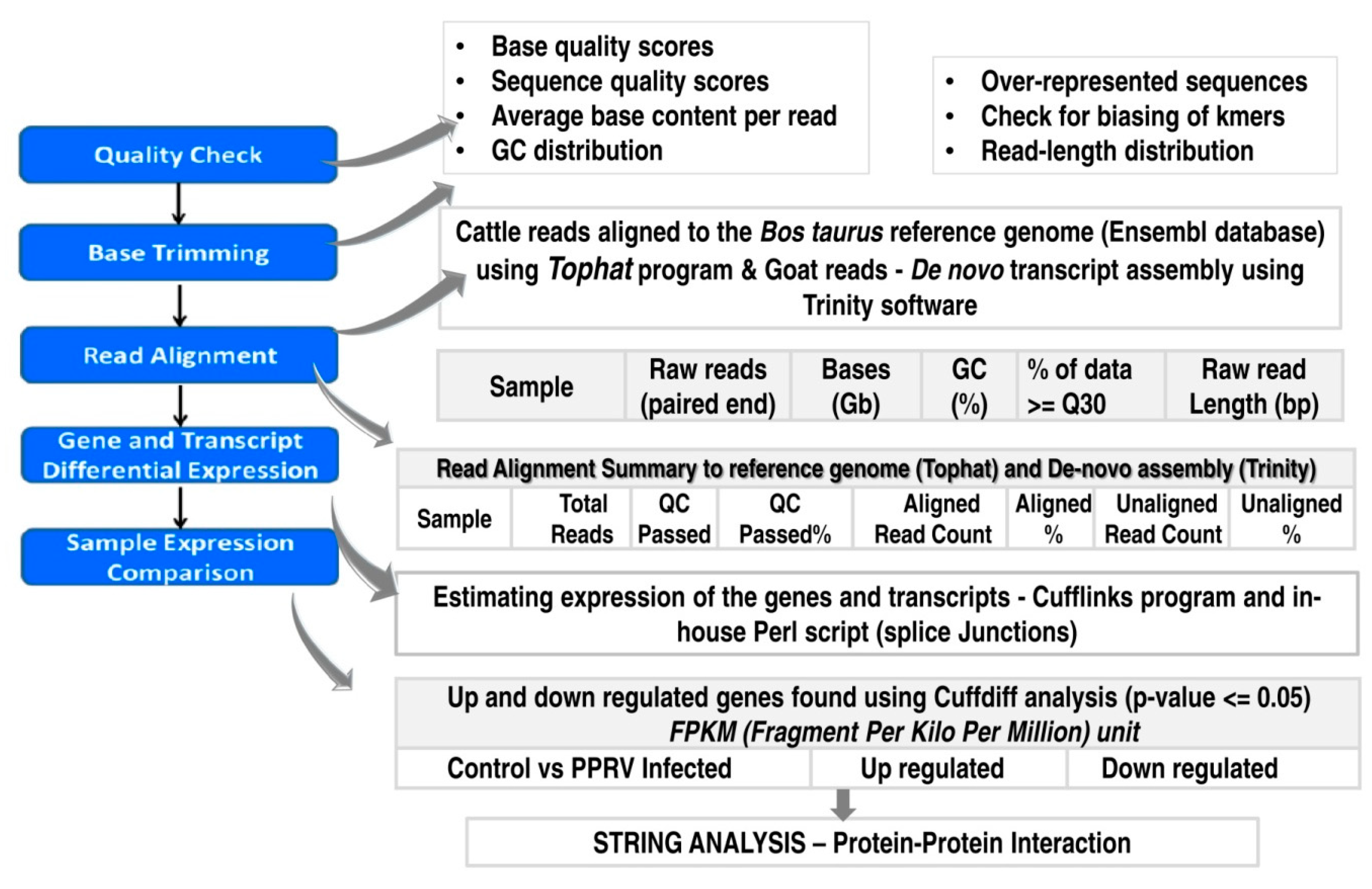
Viruses | Free Full-Text | RNAseq Reveals the Contribution of Interferon Stimulated Genes to the Increased Host Defense and Decreased PPR Viral Replication in Cattle | HTML
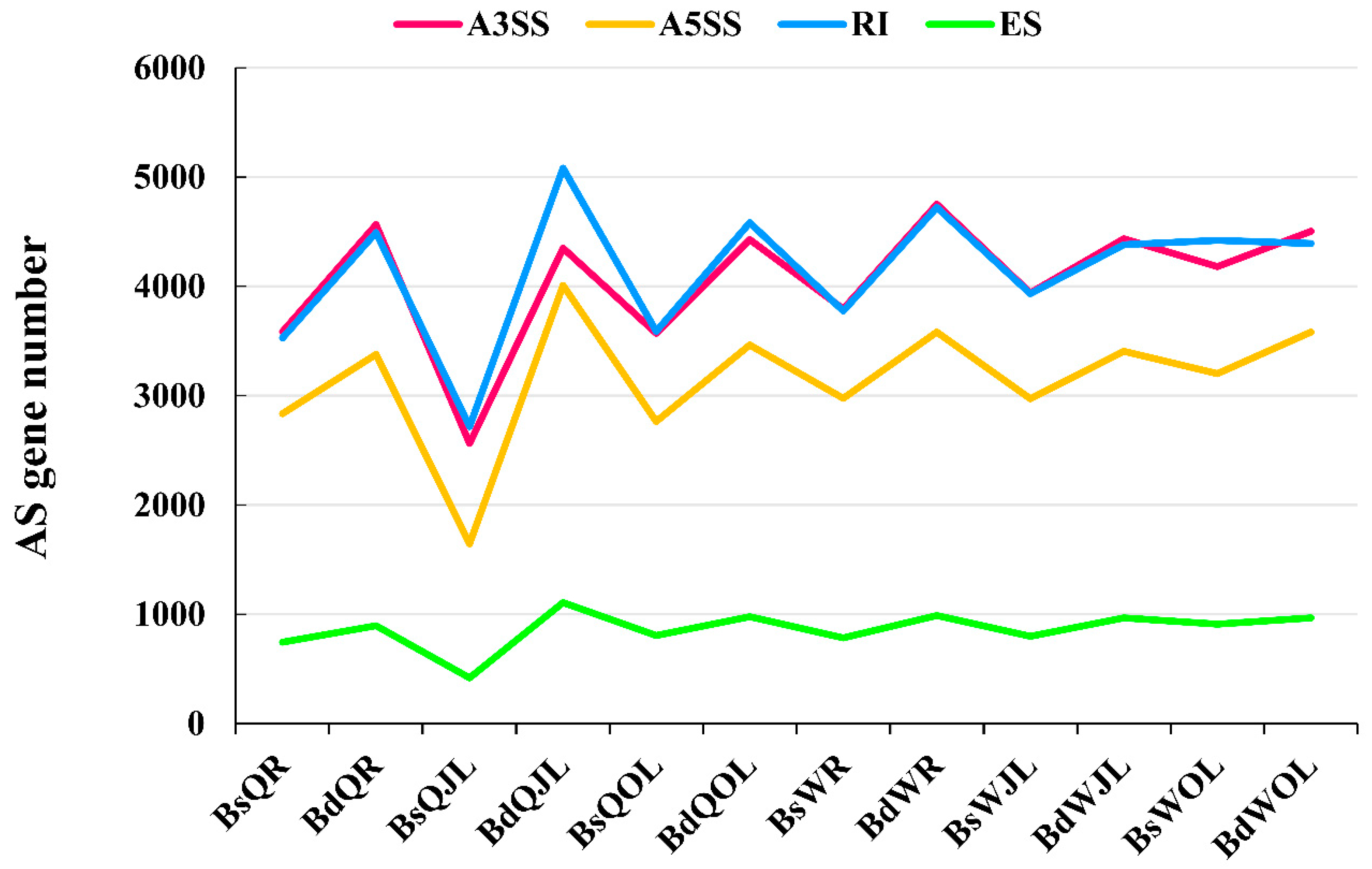
Genes | Free Full-Text | Differential Alternative Splicing Genes in Response to Boron Deficiency in Brassica napus | HTML

Sequence variation between 462 human individuals fine-tunes functional sites of RNA processing | Scientific Reports
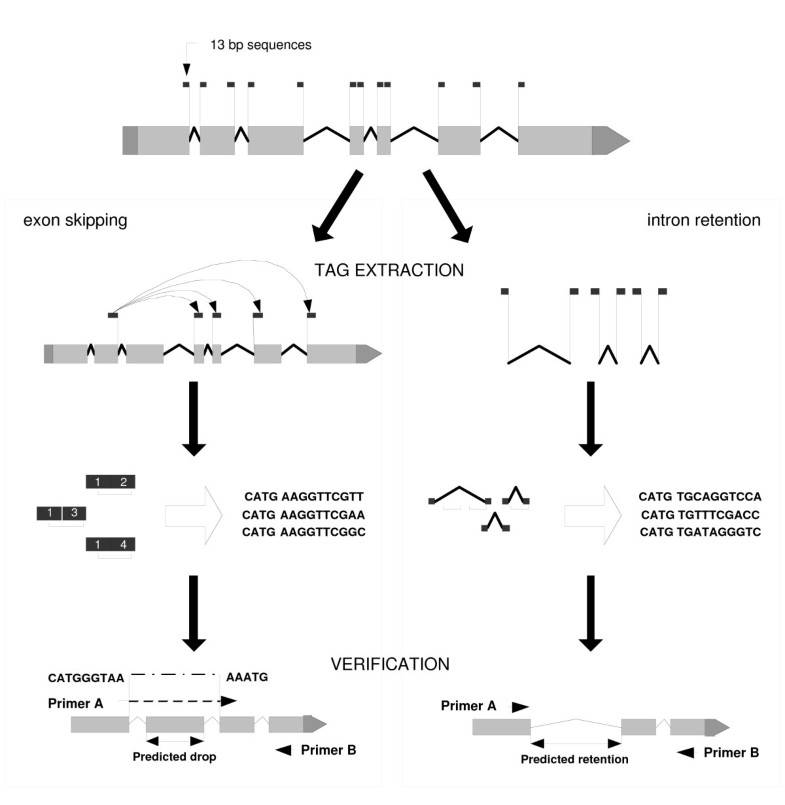
Discovery of novel alternatively spliced C. elegans transcripts by computational analysis of SAGE data | SpringerLink

IntSplice: prediction of the splicing consequences of intronic single-nucleotide variations in the human genome | Journal of Human Genetics

PDF) The Melon Zym Locus Conferring Resistance to ZYMV: High Resolution Mapping and Candidate Gene Identification

Computational approaches for circular RNA analysis - Jakobi - 2019 - WIREs RNA - Wiley Online Library
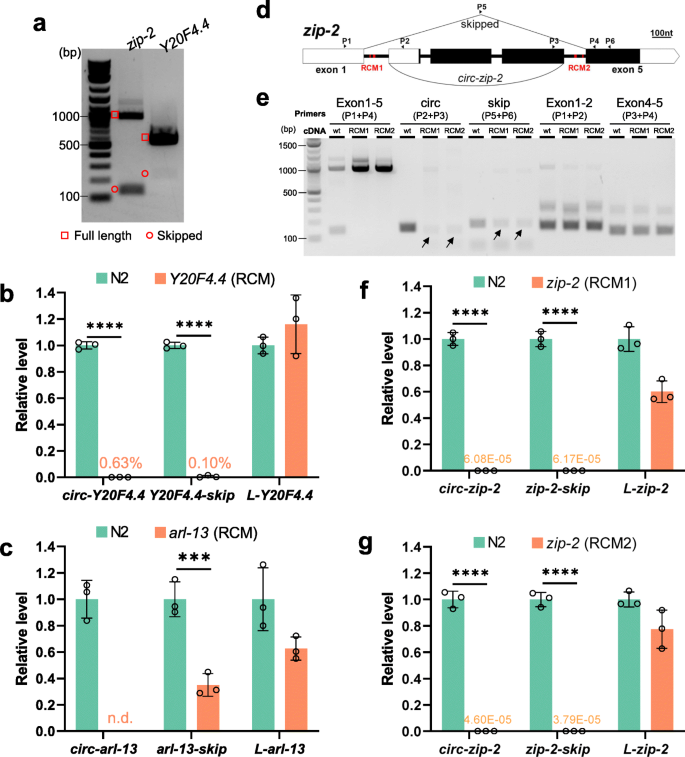
Reverse complementary matches simultaneously promote both back-splicing and exon-skipping | SpringerLink
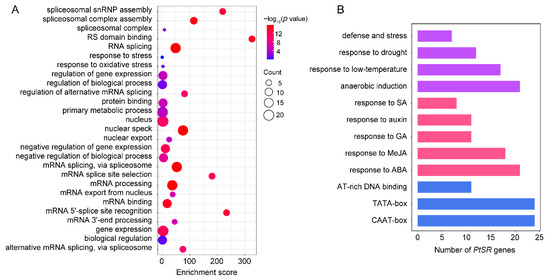
IJMS | Free Full-Text | The SR Splicing Factors: Providing Perspectives on Their Evolution, Expression, Alternative Splicing, and Function in Populus trichocarpa | HTML
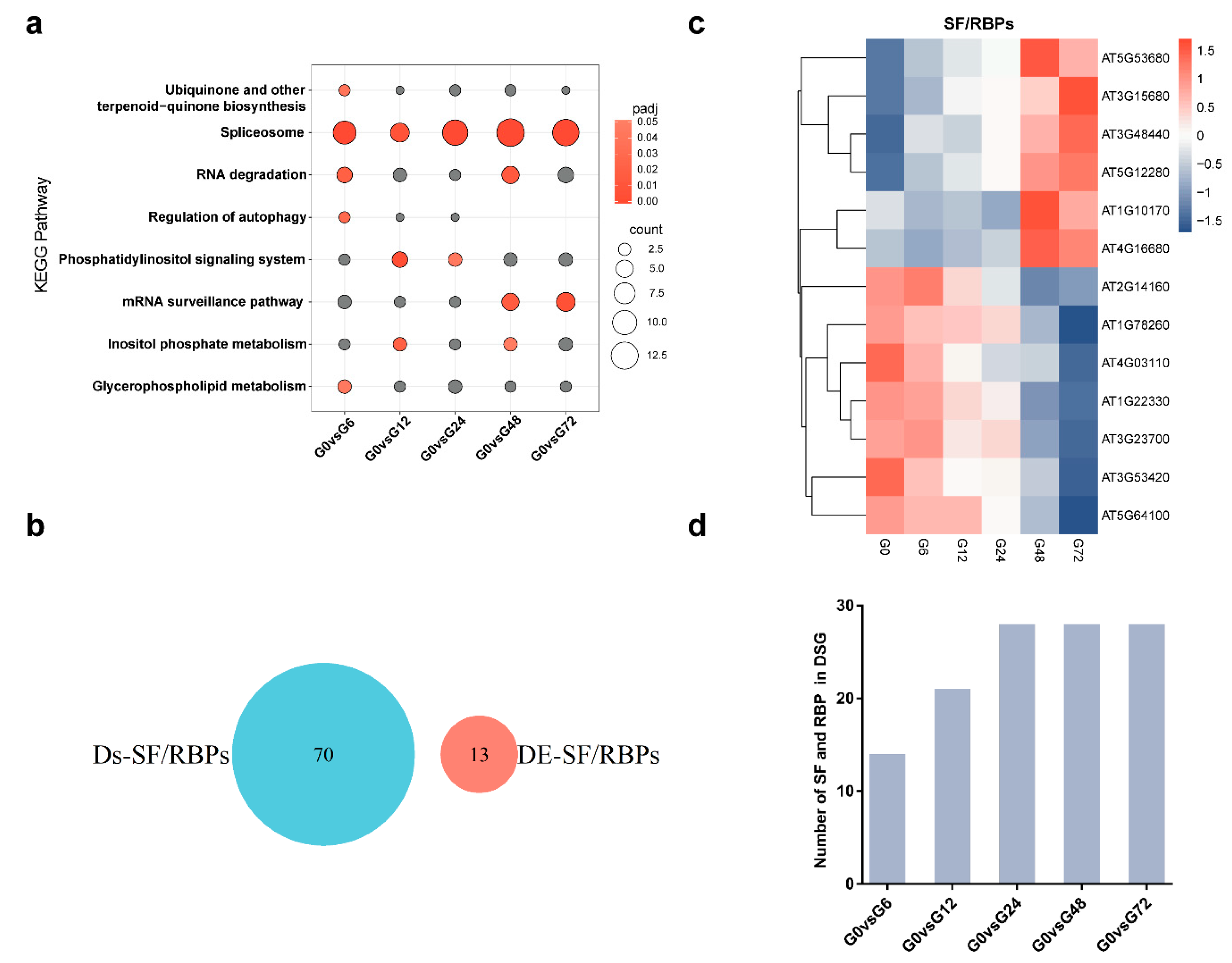
Genes | Free Full-Text | Global Profiling of Dynamic Alternative Splicing Modulation in Arabidopsis Root upon Ralstonia solanacearum Infection | HTML
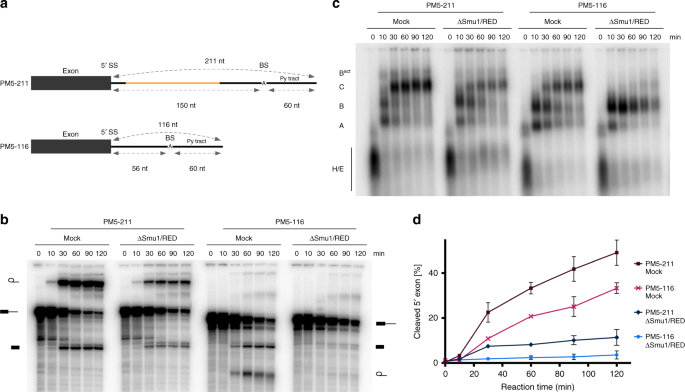
Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns | Nature Communications
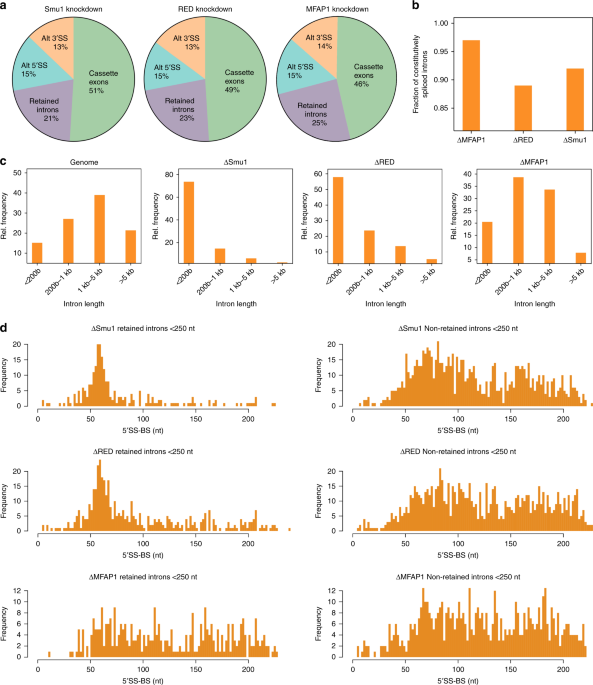
Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns | Nature Communications
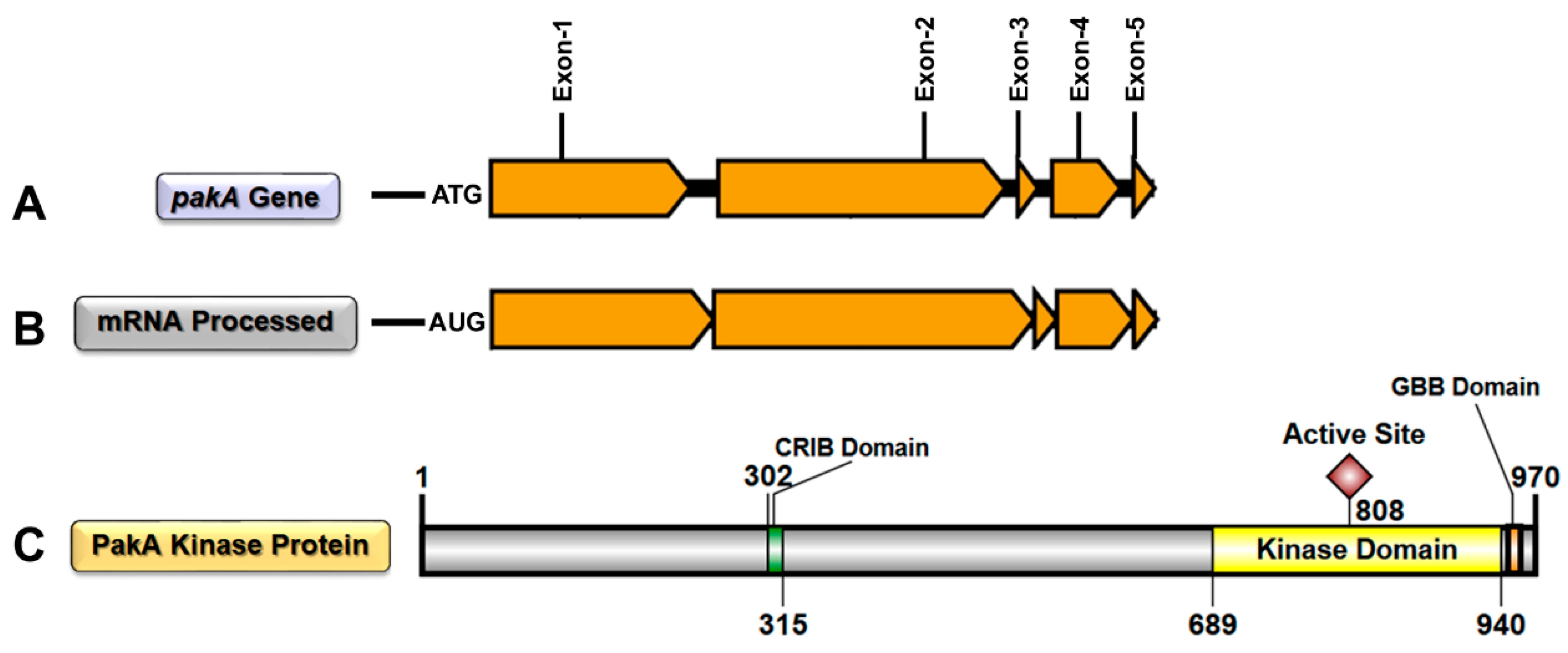
IJMS | Free Full-Text | STE20/PAKA Protein Kinase Gene Releases an Autoinhibitory Domain through Pre-mRNA Alternative Splicing in the Dermatophyte Trichophyton rubrum | HTML
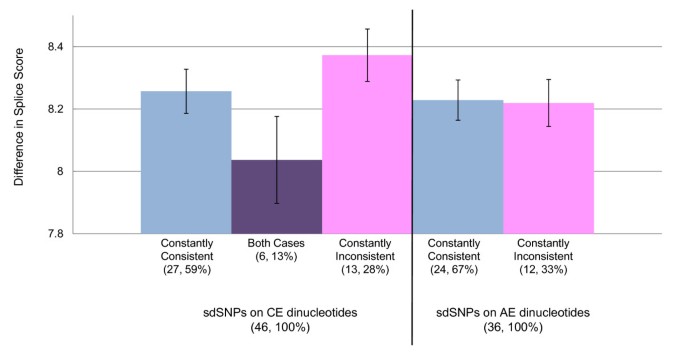
A comprehensive survey of human polymorphisms at conserved splice dinucleotides and its evolutionary relationship with alternative splicing | SpringerLink
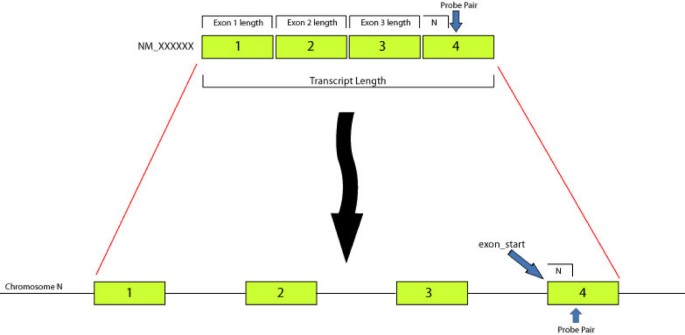
Splicy: a web-based tool for the prediction of possible alternative splicing events from Affymetrix probeset data | BMC Bioinformatics | Full Text




